[ad_1]
Disclaimer: Early release articles are not considered as final versions. Any changes will be reflected in the online version in the month the article is officially released.
Author affiliation: Applied Science, Research & Technology, Inc., Atlanta, Georgia, USA (B. Beall, S. Mathis, Z. Li, A.-K. Venero); Centers for Disease Control and Prevention, Atlanta (J. Rivers, B.J. Metcalf, L. McGee, S. Chochua)
Group A Streptococcus (GAS) commonly causes noninvasive infections affecting the skin and throat and invasive infections that can involve any tissue of the human body. Treatment of GAS infections is primarily with β-lactam antimicrobial drugs; macrolides and clindamycin are alternatives for patients allergic to β-lactam antimicrobial drugs (1). GAS co-resistance to macrolides and clindamycin has increased (2), which compromises macrolide usage for noninvasive infections and combined clindamycin with penicillin for severe disease (1). The 2 main streptococcal macrolide resistance mechanisms are 23S rRNA methylation by erm gene–encoded methylases, which confers resistance to macrolides, lincosamides (including clindamycin), and streptogramin B antimicrobials, and macrolide efflux by mef-encoded and msrD-encoded proteins (3). The lnu genes confer lincosamide resistance, whereas lsa genes confer resistance to lincosamides, streptogramin A drugs, and pleuromutilins. The pleuromutilin lefamulin is approved in the United States for systemic treatment of community-acquired bacterial pneumonia in adults (4) and has potent antibacterial activity against β-hemolytic streptococci (5). Although lsa and lnu genes are documented in group B Streptococcus (6,7) only 1 GAS isolate carrying tandem lsaE-lnuB determinants has been reported (8).
We identified 11 invasive GAS (iGAS) blood isolates positive for lsa or lnu genes, 7 lsaC and 4 lsaE/lnuB, from >16,500 iGAS isolates recovered during 2015–2023 and 335 isolates screened before 2015 through Active Bacterial Core surveillance (ABCs). We detected the positive isolates on 1 of 9 distinct mobile elements (Figures 1, 2; Appendix Table, Figures 1–9). Each element was found within 1 of 7 different iGAS strains (7 emm type/sequence type [ST] combinations; for example, emm89.0/ST101). Two strains were represented by indistinguishable (360807 and 360907) or nearly identical (20156709 and 20175626) isolate genomes obtained under BioProject PRJNA395240 (Appendix Table).
We found 5 large (61,501–78,917 bp) accessory elements carrying lsaC in combination with ermB and tetM in 6 isolates from 5 strains and with ermTR in isolate 20201216 (Figure 1). All 6 elements were flanked by short genomic target repeats, indicative of genomic insertion through precisely targeted transposition (7). Three of the 5 elements were inserted at the rplL 3′ end, 1 within the rlmD gene, and 1 within the lysS gene.
Four iGAS strains, including 1 S. equisimilis isolate (10), carried an identical lnuB allele and conserved lsaE alleles sharing 98%–100% sequence identity on 4 distinct accessory elements (Appendix Figure 2, panel B). As with 3 lsaC-carrying elements (Figure 1), 2 of the 4 elements carrying lsaE-lnuB mapped at the rplL 3′ end and were also apparently inserted through precise transposition events. For 2 strains, we were unable to map element genomic insertion sites because of incomplete assembly.
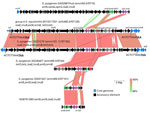
The 4 deduced 492 residue LsaE proteins shared 52.2%–53.7% sequence identity with the 5 deduced 494 residue LsaC protein sequences. Other than resistance determinants, few genes were conserved between the 4 mobile elements carrying lsaE-lnuB from this study with the prophage described from S. pyogenes strain Gas2887Hub (8) also carrying those genes (Figure 2). The iGAS S. equisimilis strain 2013217351 and S. pyogenes 20231270 carried closely related transposons, each inserted at the 12-mer rplL 3′ terminus conserved between the 2 species.
The 5 lsaC-carrying elements represented 4 phylogenetically distinct lsaC alleles (Appendix Figure 1) with 90.4%–99.8% sequence identity to the S. agalactiae UCN70 lsaC allele (6). The 4 elements carrying lsaC, ermB, and tetM each contained a small (5,258–5,816 bp) conserved lsaC self-mobilizing element inserted within a consensus Tn916 oriT site sequence (Appendix Figure 1, panel B) described in S. agalactiae (7) and recently described in 2 distinct pneumococcal elements (9). There was wide sequence divergence between the 4 small oriT-targeting iGAS lsaC mobile elements, despite identical 24–25 bp sequences flanking their insertion sites. For the lsaC-containing element in strain 20156709/20175626, genomic insertion and phylogenetic data were consistent with the sequential genomic insertion of a Tn916 family element before a more recent second precise insertion of the 5546 bp lsaC–carrying element into its oriT site (Appendix Figure 2, panels A, B). For 2 other lsaC-carrying elements, phylogenetic data suggested recent introduction of the complete composite element, consisting of a Tn916-related element carrying an integrated small lsaC element (Appendix Figures 3–4).
Ten of the 11 study isolates were resistant to both erythromycin and clindamycin (Appendix Table); that resistance is associated with the presence of ermB and lsaC (6 isolates), ermTR and lsaC (1 isolate), or ermB, lsaE, and lnuB (3 isolates). One strain, 20201347 (lsaE+, lnuB+), was erythromycin susceptible but clindamycin-resistant, indicating lsaE- and lnuB-conferred clindamycin resistance. That finding in strain 20201347 was consistent with masking of lsaE- and lnuB-conferred clindamycin resistance in the other 10 isolates because of erm gene–encoded methylase activity. The 4 isolates carrying lsaE-lnuB had high MICs for the pleuromutilin lefamulin (MIC >2 µg/mL), whereas the 7 lsaC-positive isolates had low MICs for lefamulin (MICs 0.25 µg/mL) (Appendix Table). We conclude that expansion of lsaE-positive iGAS lineages could compromise future potential use of lefamulin, and lsaC– or lsaE-lnuB–positive strains could further undermine the use of clindamycin for treating β-hemolytic streptococcal infections.
Dr. Beall has been conducting streptococcal strain surveillance and ABCs-related streptococcal research since 1994. After retiring from the National Center for Immunization and Respiratory Diseases, Centers for Disease Control and Prevention, in 2021, he continues to work on ABCs-related streptococcal surveillance projects.
[ad_2]
Source link